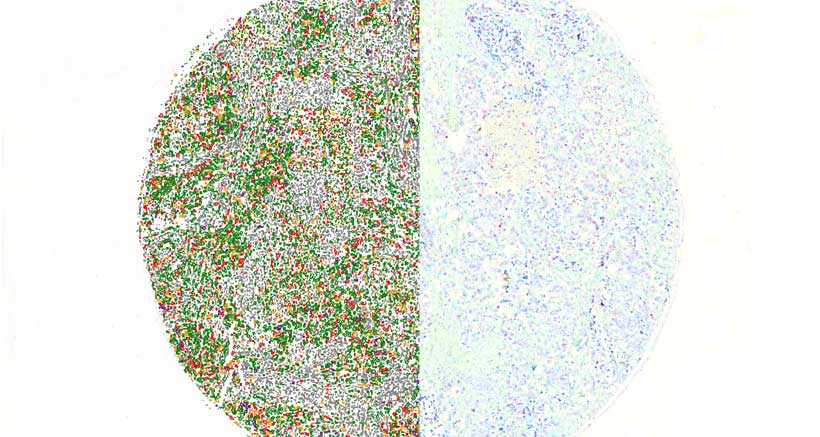
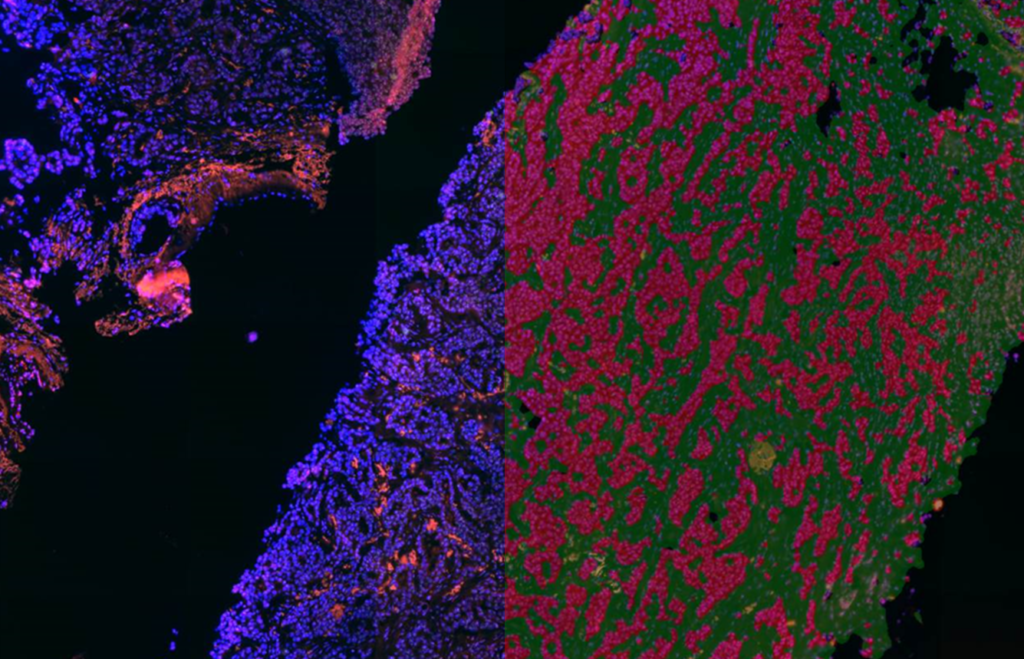
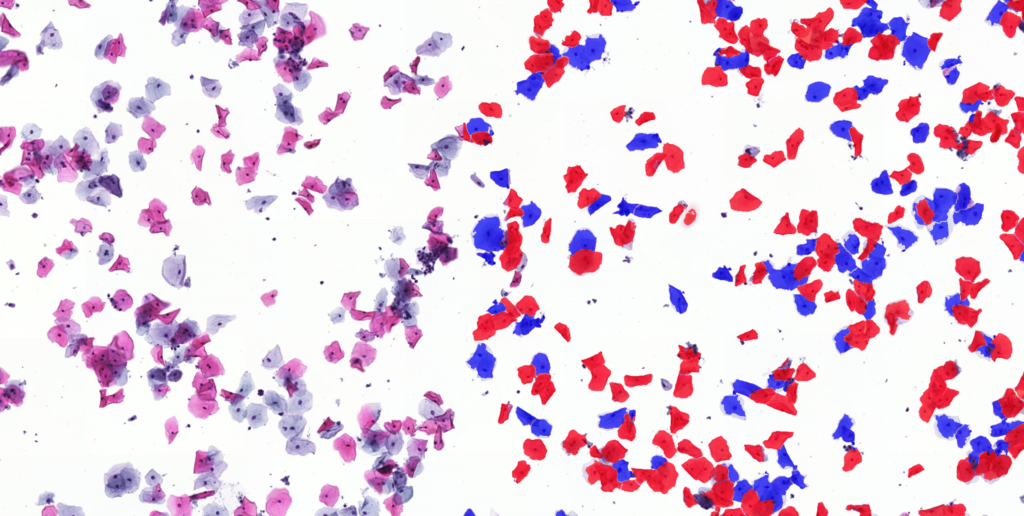
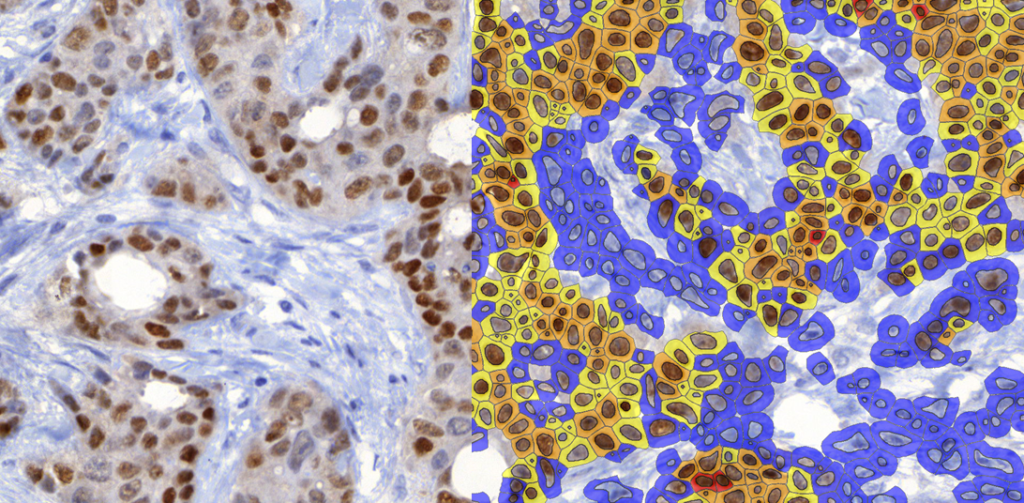
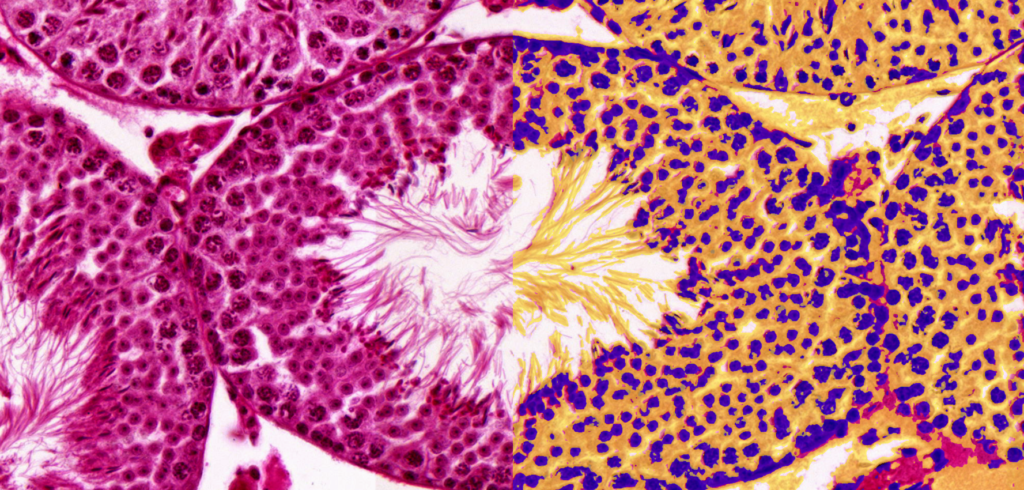
QuantCenter is an image analysis platform for whole-slide quantification in histopathology and molecular pathology. With customizable algorithms and linkable modules. Integrated with SlideManager, it enables efficient execution of analysis on research study samples, driving precise reproducible results. It offers a range of combinable modules including:
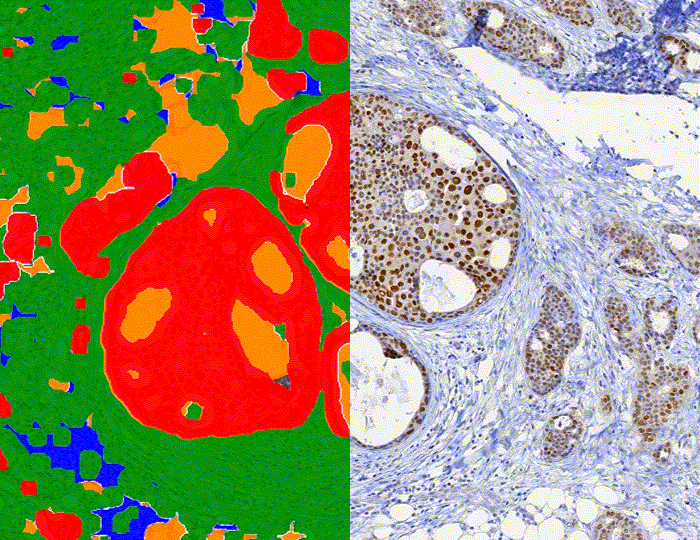
PatternQuant
A trainable pattern recognition module for tissue classification and pre-segmentation.
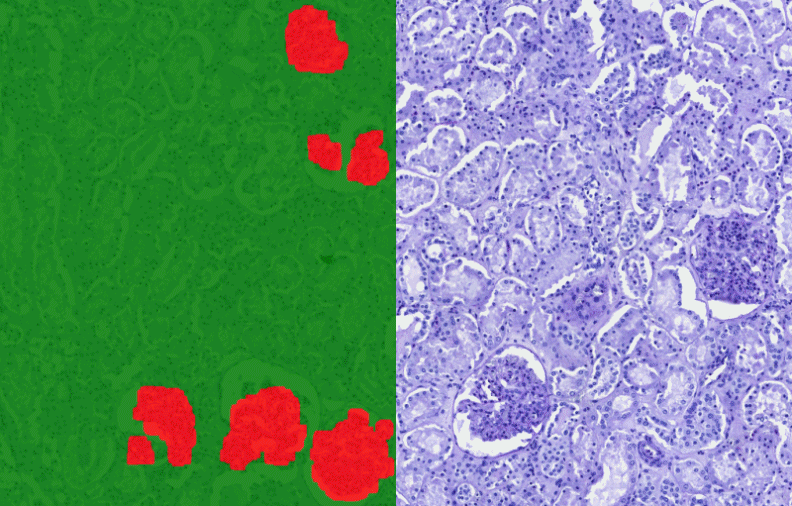
PatternQuant Plus
An enhanced version of PatternQuant, utilizing deep learning for complex tissue segmentation projects.
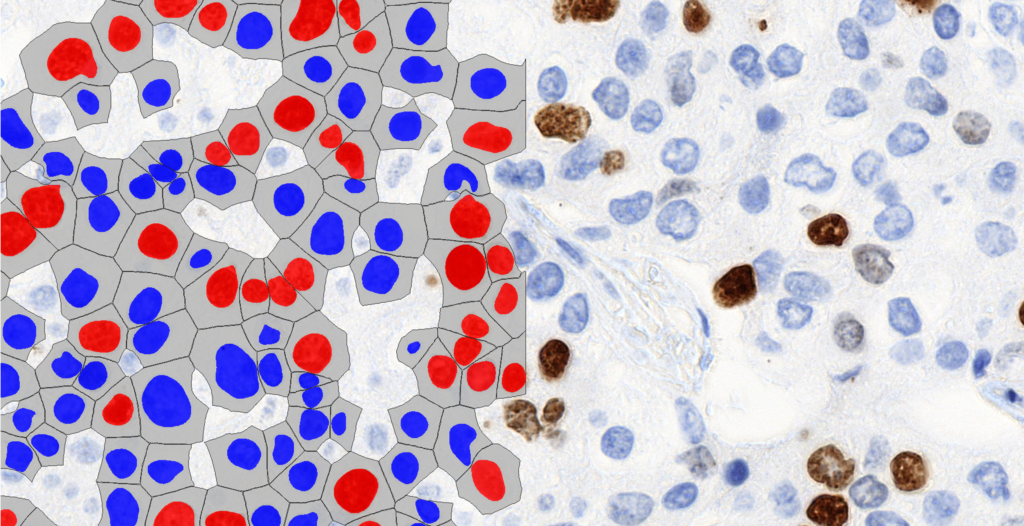
CellQuant
A versatile cell detection application suitable for various IHC stainings, including Ki67 slides.
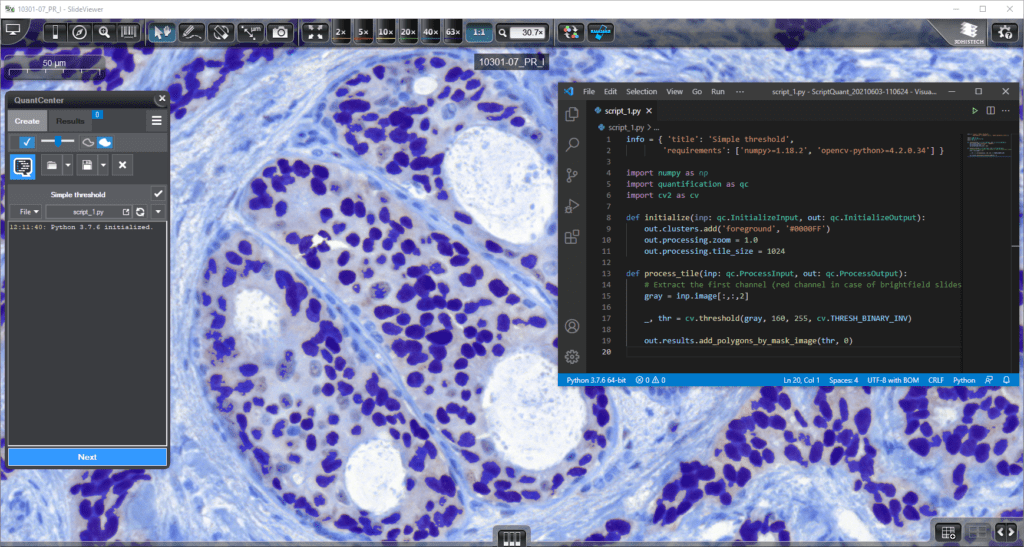
ScriptQuant
Allows users to run custom image analysis algorithms within the QuantCenter framework.
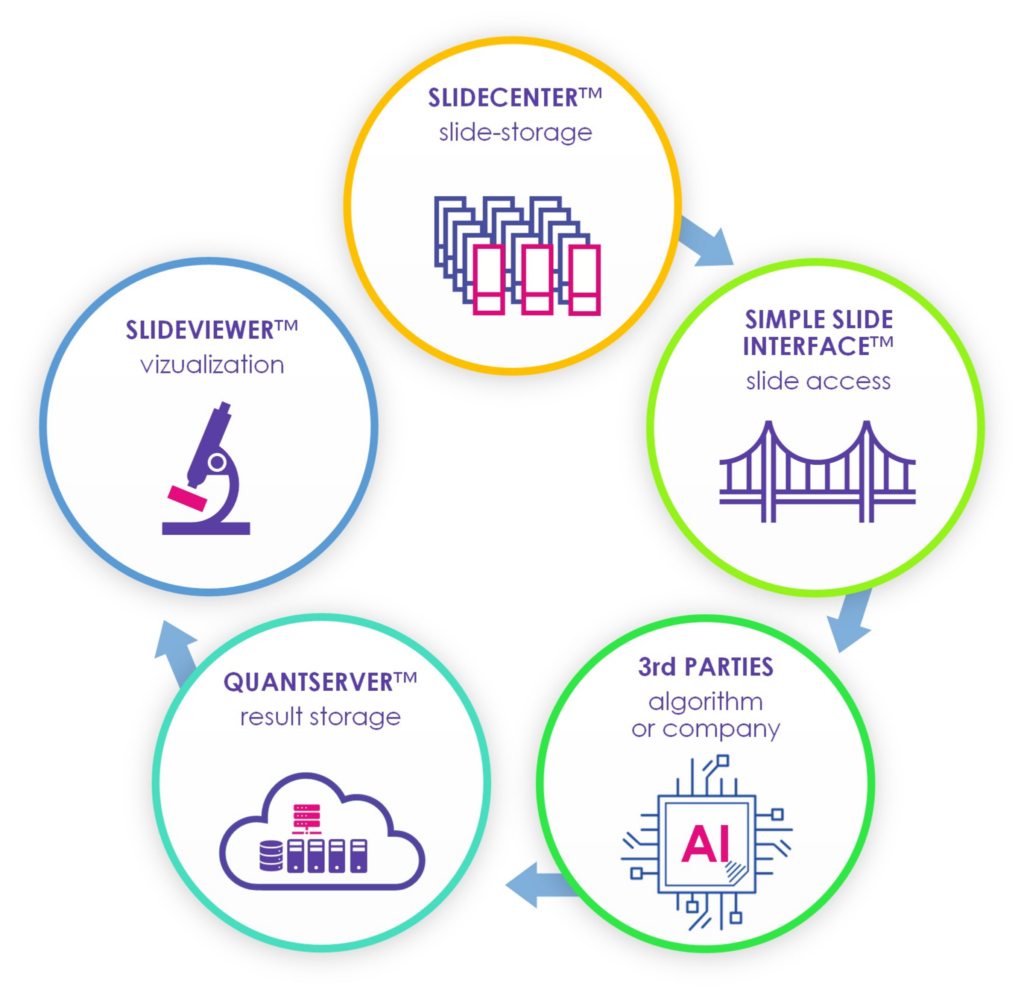
SimpleSlideInterface
Enables easy access to slide-related data for custom application development.
Additional quantification tools include:
BatchAnalysis-ProcessingQueue
Allows multiple digital slides to be examined in the background, saving time.
Data Visualization (DVT)
Provides integrated data visualization modes, including tables, scatterplots, pie charts, and histograms.
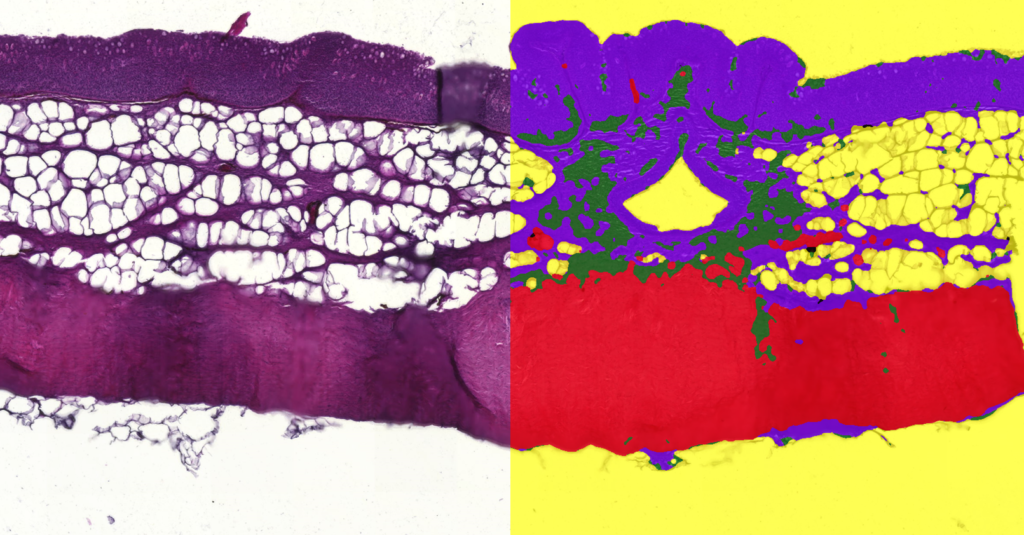
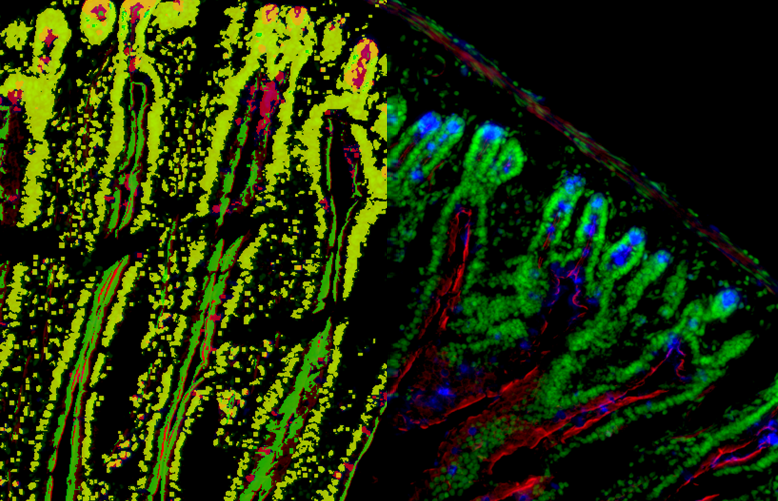
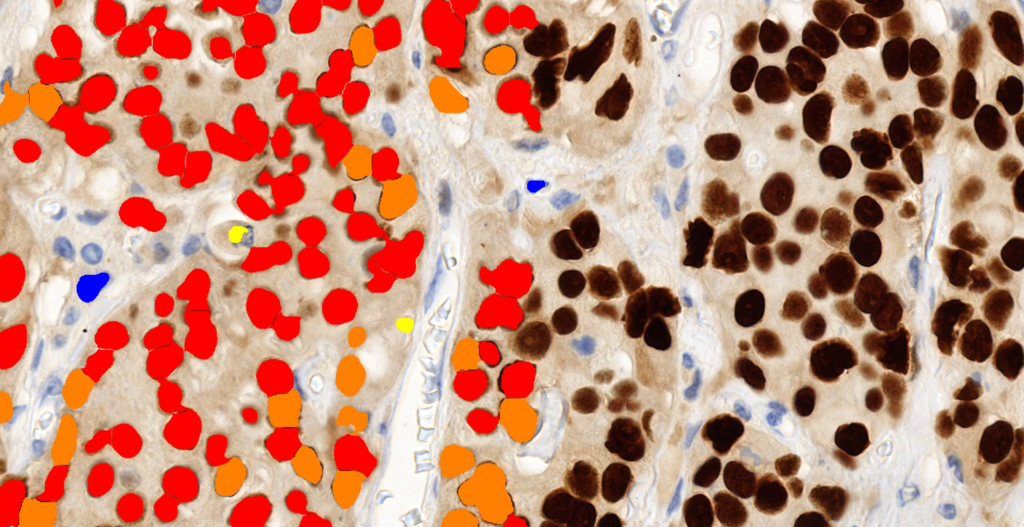
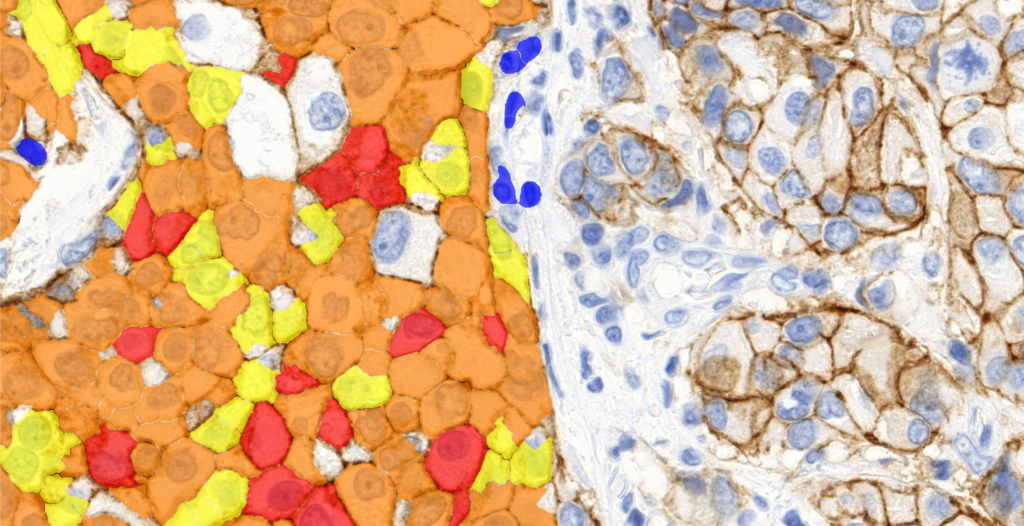
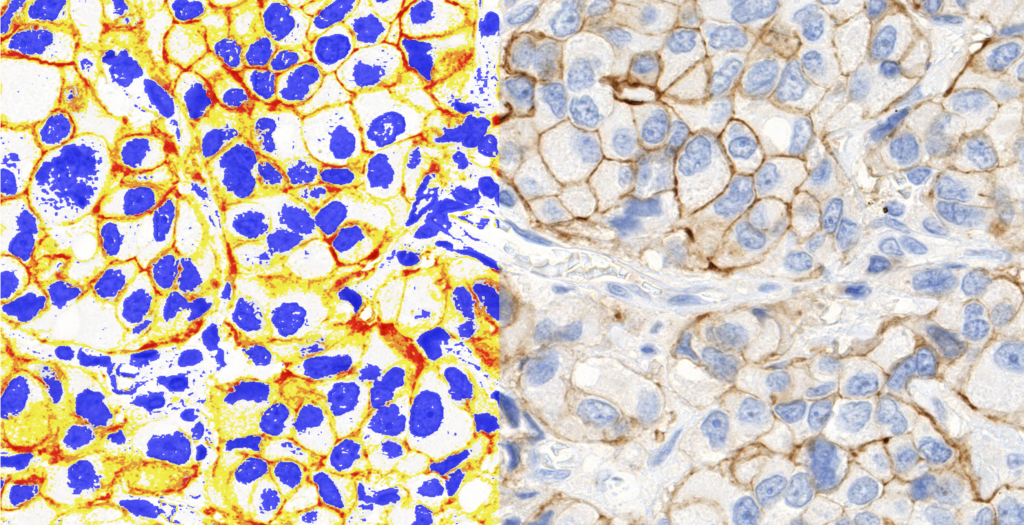
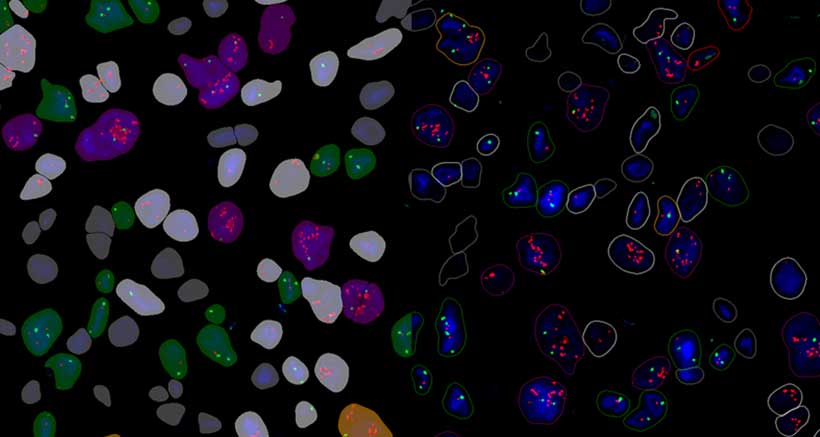
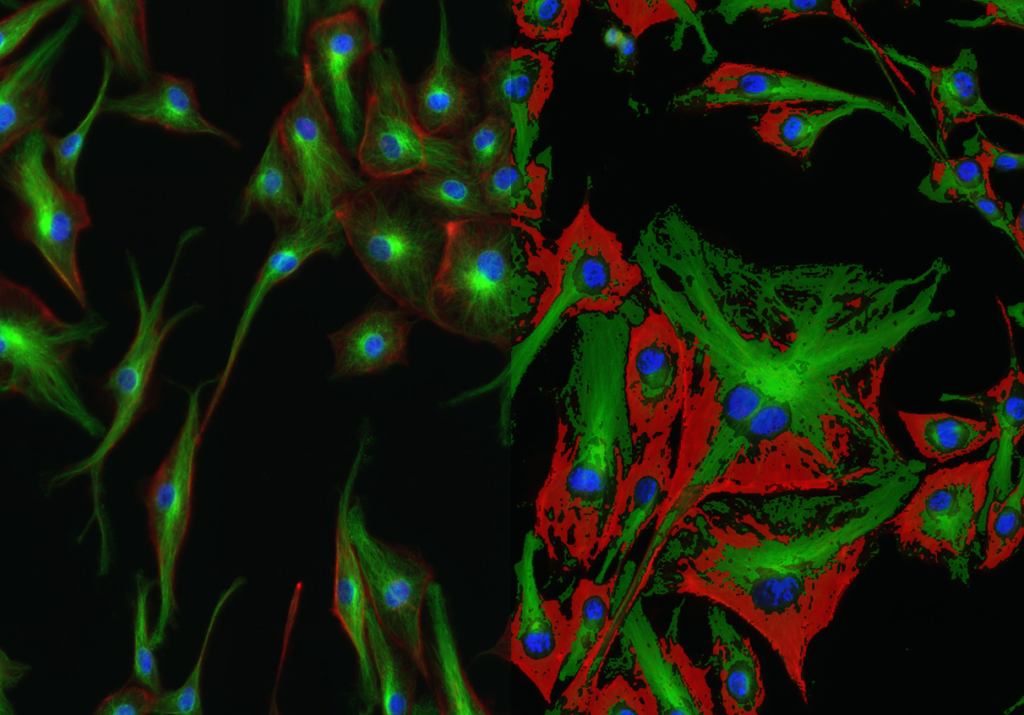
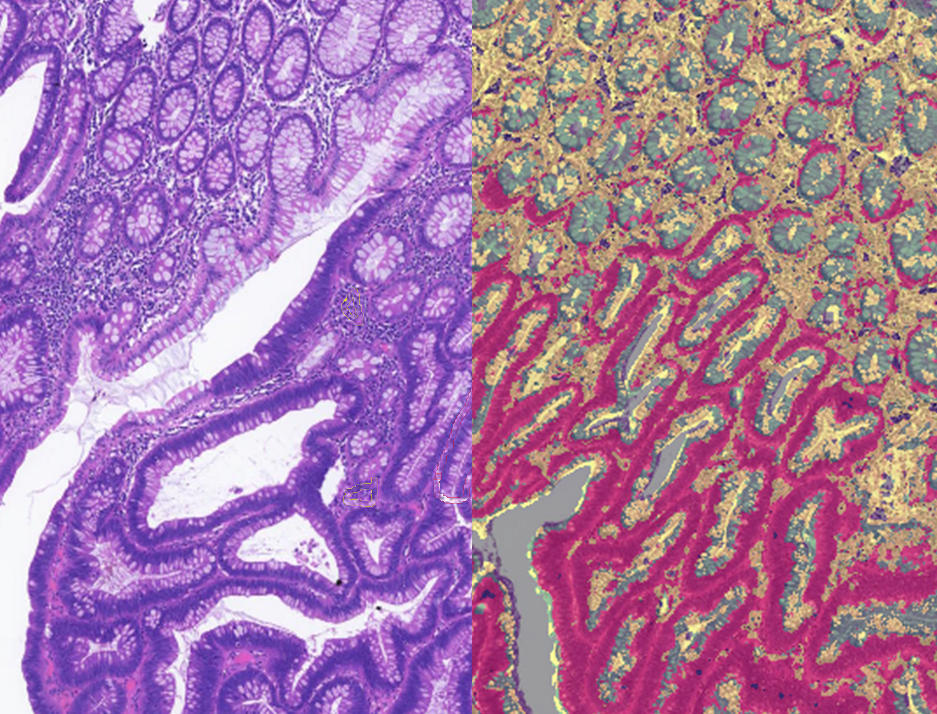
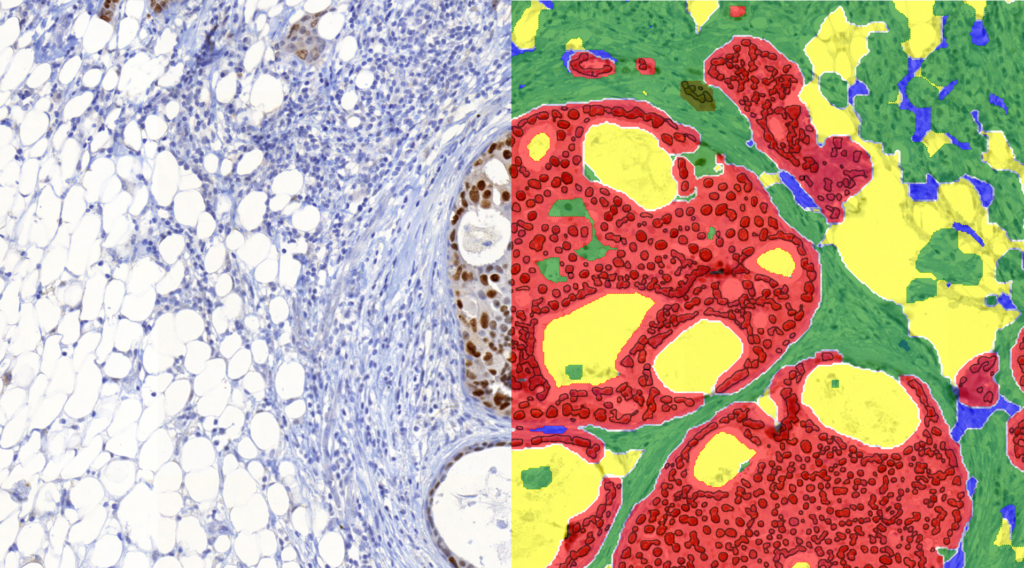
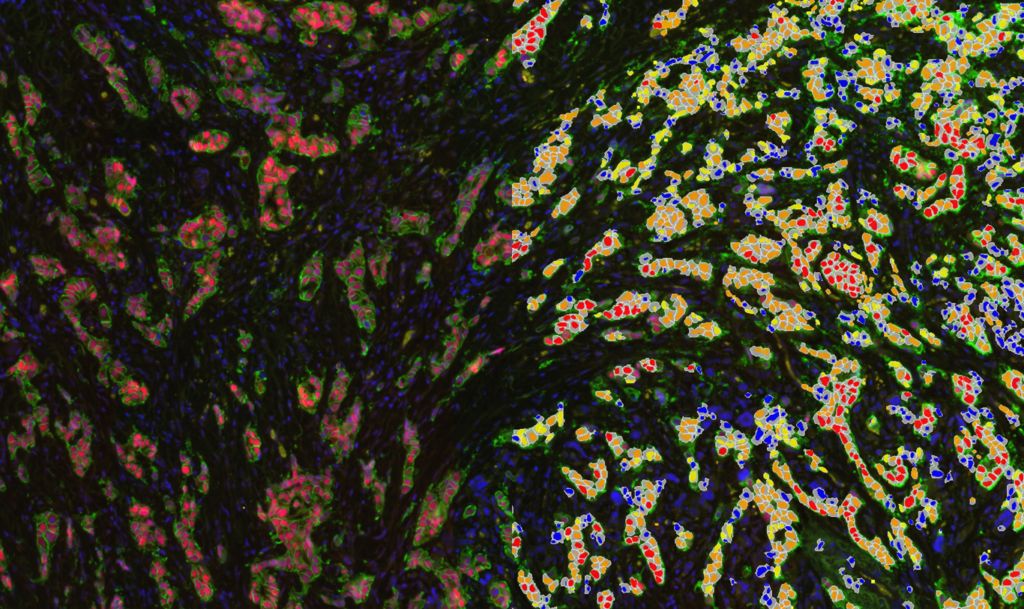
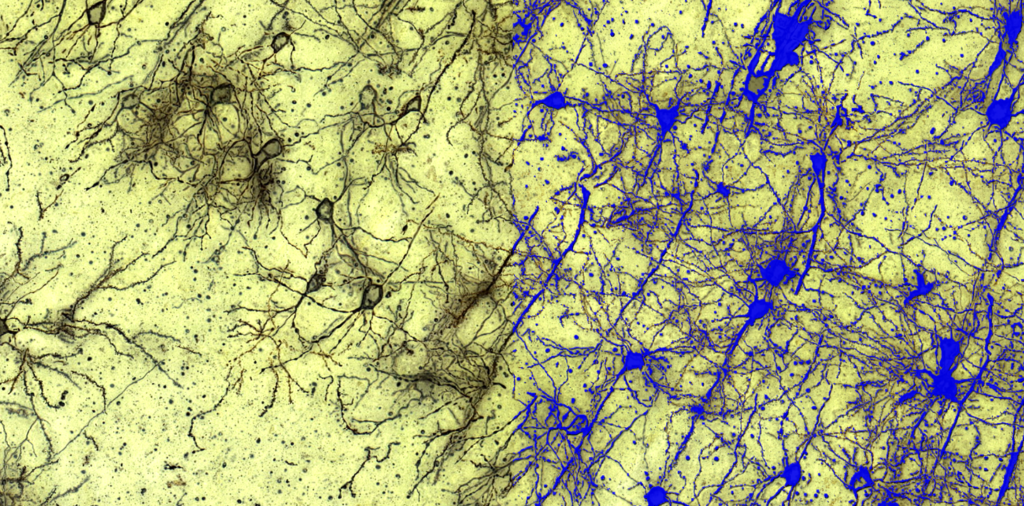
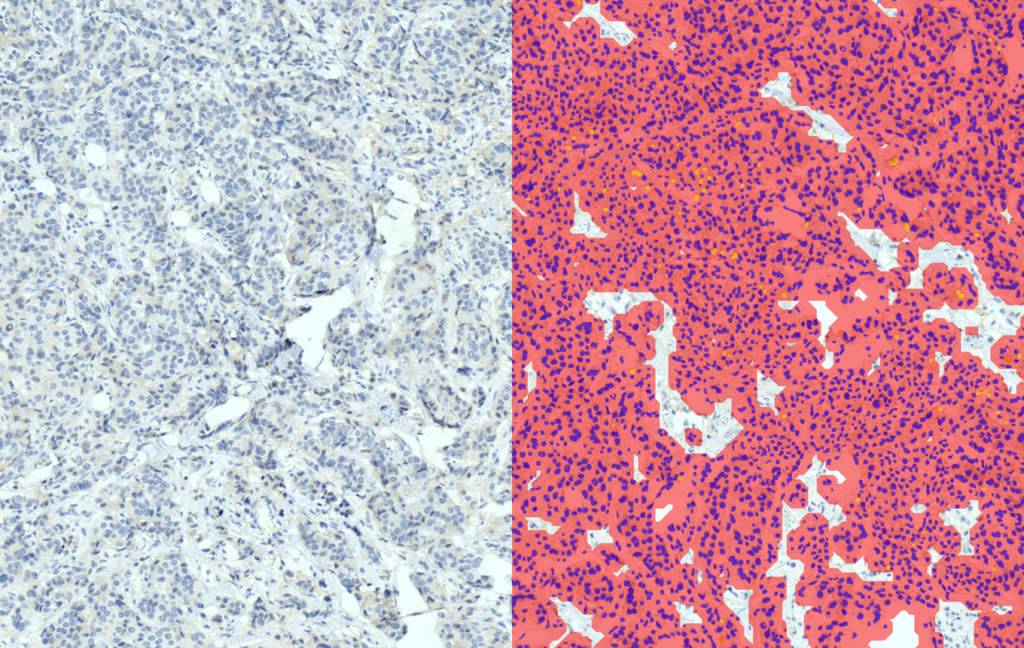
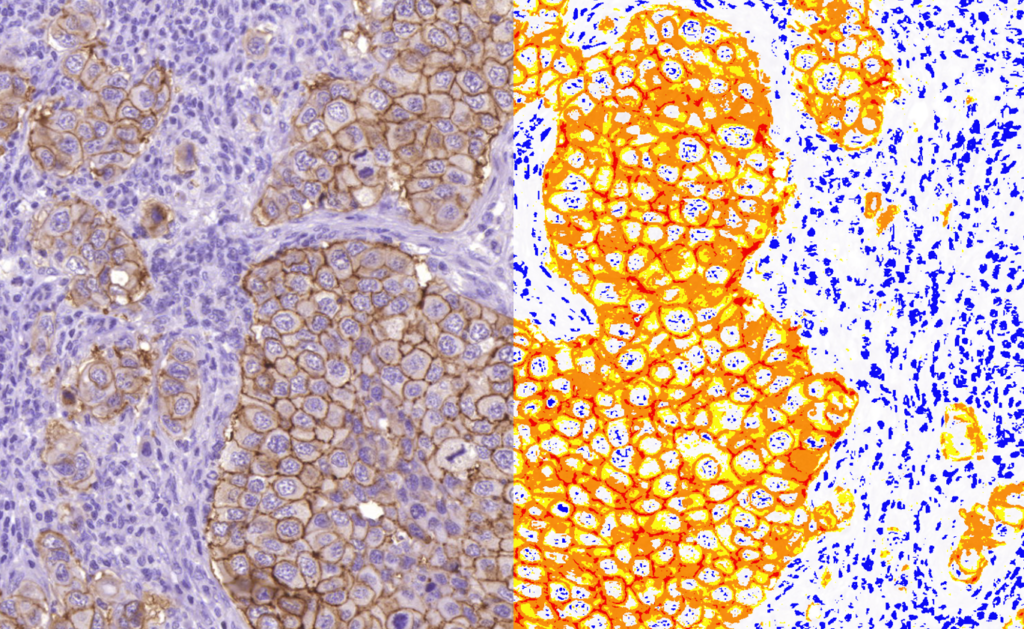
Quantification Examples
The gallery below contains some examples of how widely our quantification modules can be used.
Click on the image to view in details.
Key features
Modular and Customizable Image Analysis
Unlike traditional image analysis software, QuantCenter offers a modular framework, allowing users to select and combine specialized tools for precise whole-slide quantification.
Customizable AI-driven analysis enables adaptation to specific research needs, providing greater flexibility than static competitor solutions.
Comprehensive Range of Analysis Modules
Advanced segmentation and classification tools, including HistoQuant, PatternQuant, and PatternQuant Plus, ensure accurate stain identification and deep-learning-driven tissue recognition.
Targeted quantification with NuclearQuant, MembraneQuant, CellQuant, DensitoQuant, FISHQuant, and CISHQuant, covering all major staining techniques.
Custom script integration via ScriptQuant, offering unmatched flexibility in digital pathology.
Superior Performance and Batch Processing
BatchAnalysis-ProcessingQueue allows multiple digital slides to be processed simultaneously, increasing efficiency and throughput.
QuantServer ensures high-speed processing and centralized data storage, optimizing workflows for large-scale research studies.
Advanced Data Visualization for Better Insights
Integrated Data Visualization (DVT) tools, including scatterplots, pie charts, histograms, and tables, enable intuitive and real-time result interpretation.
Quantitative analysis results can be easily exported for reporting and collaboration.
Seamless Integration with Digital Pathology Workflows
Fully compatible with SlideManager, ensuring smooth execution of analysis
on study samples without requiring additional manual steps.
Supports a wide range of whole-slide image formats, making it a universal solution for diverse pathology laboratories.
Scalable for Research and Clinical Applications
Suitable for small-scale research and high-throughput clinical studies, with adaptable computational power and server-side processing.
Enables standardized, reproducible image quantification, essential for clinical diagnostics and drug development.